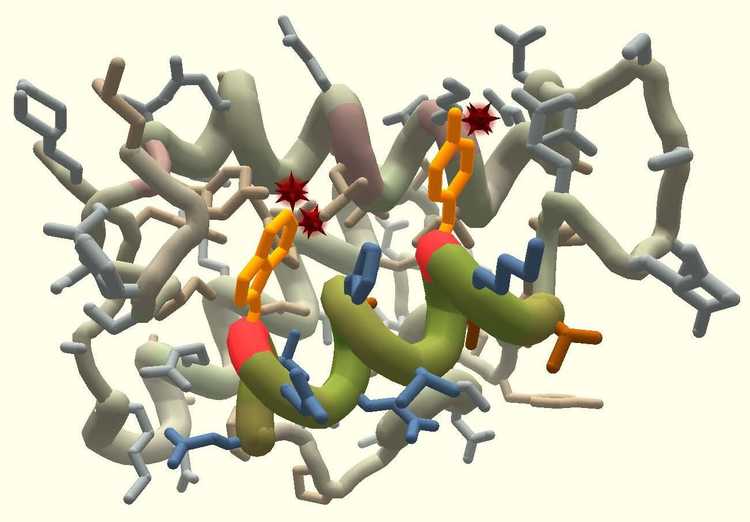
Foldit indirectly challenges gamers to predict and design the structures of proteins in the hopes of better understanding how they work and why they might be causing trouble in the human body. “With all the things proteins do to keep our bodies functioning and healthy, they can be involved in disease in many different ways. The more we know about [them], the better new proteins we can design to combat the disease-related proteins and cure the diseases,” project lead Seth Cooper explains.
Playing Tetris to advance biochemistry
Foldit attempts to predict the structure of a protein by taking advantage of humans’ puzzle-solving intuitions and has people play competitively to fold the best proteins. “The approach we have taken is to combine the spatial reasoning and problem solving of people with the number-crunching optimisation power of computers,” Popovic adds. Thousands of people around the world have played Foldit already—a game much like the tile-matching puzzle video game Tetris—to help biochemists to tackle protein-related questions.
Remarkably, few of the gamers that are now actively involved have a background in biochemistry. Yet still, they are working to design brand new proteins that could help prevent or treat diseases such as HIV/AIDS, cancer, and the flu. “There are all kinds of players of different professions, ages, and nationalities.” The fact that people can play the game together has proven to be a very important feature of the game. “Players can form groups and share the solutions that they have found. Players may have different skills, so by working together they may be able to find things that they wouldn’t have found alone.”
Protein folding by mass collaboration and creativity
The game helps scientists to gain insight into the “behaviour” of proteins. “Proteins are the workhorses in every cell of every living thing,” Cooper explains. “Your body is made up of trillions of cells, of all different kinds: muscle cells, brain cells, blood cells, and more. Inside those cells, proteins are allowing your body to do what it does: break down food to power your muscles, send signals through your brain that control the body, and transport nutrients through your blood.” When proteins stop working properly, disease occurs.
Essentially, a protein is made up of a long chain of amino acids. To make a protein, the amino acids form an unbranched chain, much like a line of people holding hands. Depending on the specific make-up of the protein, the chain folds up into a very specific shape—the same shape every time. “Most proteins do this all by themselves, although some need extra help to fold into the right shape. The unique shape of a particular protein is the most stable state it can adopt.”
While gaming, research groups take the lead and define sub-challenges with specific goals and frameworks, carrying names like 675: Hydrogen-producing Catalyst Dimer. Players can be individuals or groups, such as l’Alliance Francophone, Team Hungary and Void Crushers. Introductory levels teach the rules, which are the same laws of physics by which protein strands curl and twist into three-dimensional shapes. These are key to solving biological mysteries ranging from Alzheimer's to vaccines.
Teaching human strategies to computers
The number of different ways in which a protein can fold is astronomical, because there are so many degrees of freedom. Figuring out which of the many possible structures is the best one is considered one of the hardest problems in biology today. Foldit accommodates this problem: “We’ve already found that for certain types of structural rearrangement of partially folded proteins, players were often able to find better solutions than purely algorithmic approaches. Players were also able to resolve the structure of a virus protein that had remained unsolved by any experimental or computational method. Experimentalists had been working on it for over ten years.” Foldit uses the newly acquired knowledge to start teaching human strategies to computers, so that they will ultimately become able to fold proteins faster than ever before.